The "Weighted Burgers Vector" is a method to analyze geometrically necessary dislocations from EBSD maps, complementary (or alternatively) to GND analysis. The reference description is in Wheeler et. al., The weighted Burgers vector: a new quantity for constraining dislocation densities and types using electron backscatter diffraction on 2D sections through crystalline materials
The "Weighted Burgers Vector" is a moving window technique interpreting the orientation gradient in a given neighborhood and at a given distance to be caused by dislocation lines which crosscut the observation surface. It is weighted/biased by the cosine of the angle between the map-normal the dislocation line direction.
In MTEX the weighted Burgers vector is computed by the function ebsd.weightedBurgersVec using one of the following two methods. The default method is derived from the orientation gradient in x- and y- directions, obtained using a convolution kernel similar to a Prewitt operator, except that the corners are weighted by 0.5. This is comparable to what is termed the "integral method" by other implementations. Here the size of kernel can be adjusted in order to cope with noisy data. Alternatively, there is the "gradient" method, where the WBV is derived from the last column of the dislocation density tensor. Hence, using high angular resolution EBSD data or denoising of conventional EBSD prior to the analysis is required.
We start by importing the same data as for the GND example.
% import the EBSD data
% mtexdata single
CS = crystalSymmetry('Fm3m',[4.04958 4.04958 4.04958],'mineral','Al');
ebsd = EBSD.load([mtexDataPath filesep 'EBSD' filesep 'single_grain_aluminum.txt'],...
'CS', CS,'RADIANS','ColumnNames', { 'Euler 1' 'Euler 2' 'Euler 3' 'x' 'y'},...
'Columns', [1 2 3 4 5]);We reconstruct grains because later on, we do not want to compute the WBV across grain boundaries
[grains,ebsd.grainId] = calcGrains(ebsd,'angle',2.5*degree,'minPixel',6);
% denoise the data
% we will use the noisy data later on
ebsdN = ebsd.gridify;
F = halfQuadraticFilter;
ebsd = smooth(ebsd('indexed'),F,'fill',grains);Computing the WBV
The function expects the EBSD data set to be gridified.
ebsd = ebsd.gridify;
wbv = weightedBurgersVec(ebsd)wbv = vector3d (y↑→x)
size: 101 x 101The WBV is returned in specimen coordinates as a list of vector3d. We can inspect its magnitude (in 1/scanunit) and direction.
plot(ebsd,wbv.norm)
mtexColorbar
mtexTitle('norm of the WBV')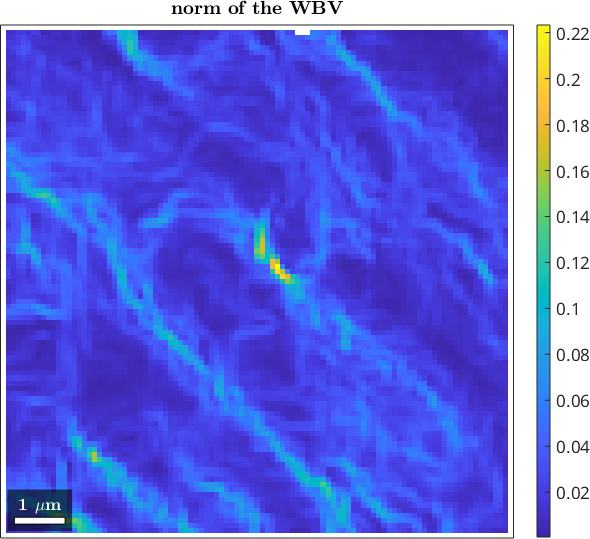
Visualizing the WBV
In order to visualize the direction of the WBV in specimen coordinates, we can use a directional color key.
cK = HSVDirectionKey;
plot(ebsd,cK.direction2color(wbv),'FaceAlpha',wbv.norm/0.22)
mtexTitle('direction of the WBV in specimen coordinates')
plot(cK,'figSize','tiny')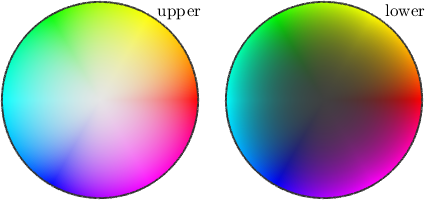
We could also display the WBV as small arrows. If we allow for any magnitude, the plot would become quite cluttered. Hence, we will only display those vectors which have a reasonably large magnitude.
cond = wbv.norm > quantile(wbv.norm,0.85);
plot(ebsd,cK.direction2color(wbv),'FaceAlpha',wbv.norm/0.22)
hold on
quiver(ebsd(cond),wbv(cond),'color','k','autoScaleFactor', 2, 'antipodal');
hold off
If we are simply interested in the distributions of WBV, we can plot them in a spherical projection. Since the WBV is assigned nan at points where it cannot be defined, we need to filter those values
wbv.antipodal = 0;
notNan = ~isnan(wbv);
plot(wbv(notNan),'weights',wbv(notNan).norm,'contourf')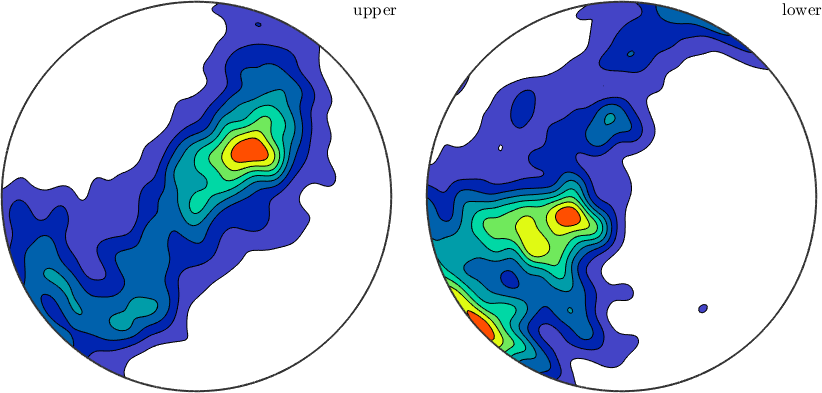
The WBV in crystal coordinates
In order to inspect the WBV in crystal coordinates we transform them using the orientations and choose a fitting directional color key
% transform to crystal reference frame
wbvC = inv(ebsd.orientations) .* wbv;
% define a directional color key respecting crystal symmetry
cKC = HSVDirectionKey(wbvC);
% plot the data
plot(ebsd,cKC.direction2color(wbvC),'FaceAlpha',wbv.norm/0.22)
mtexTitle('direction of the WBV in crystal coordinates')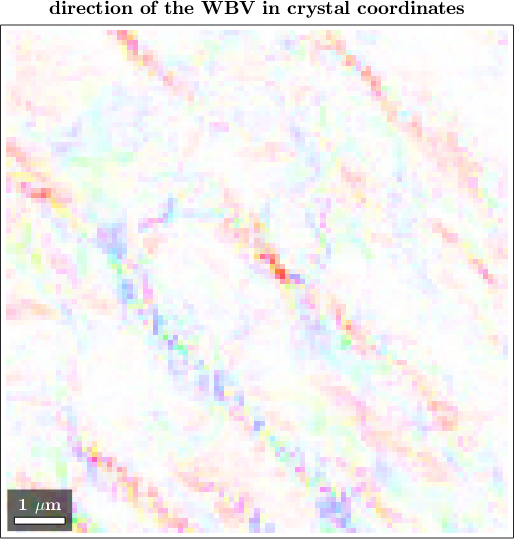
% plot the color key
plot(cKC)
% overlayed with the
hold on
plot(wbvC(notNan),'weights',wbvC(notNan).norm,'contour', ...
'contours',0.5:0.25:2,'linecolor','k','ShowText','on', ...
'linewidth',2)
hold off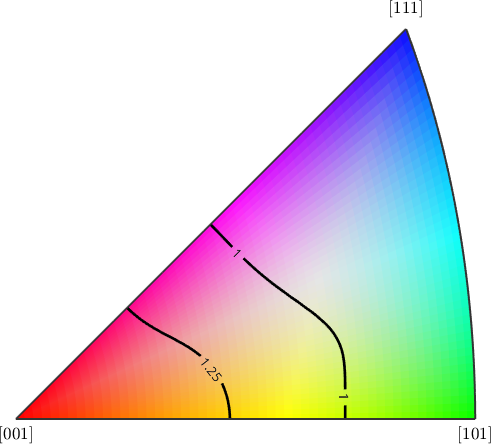
Effect of windowSize
In case there are reasons why the EBSD data cannot be denoised, the WBV can also be computed with respect to a larger neighborhood. The integer specified with with 'windowSize' gives a 2*n+1 square across which the WBV is computed. The default is a 3-by-3 box.
close all
newMtexFigure('layout',[2,4])
% first we plot again the WBV form the denoised dataset
wbv = weightedBurgersVec(ebsd);
nextAxis(1,1)
plot(ebsd,wbv.norm); hold on
mtexTitle('norm (denoised) / box = 3')
nextAxis(2,1)
notNan = ~isnan(wbv);
plot(wbv(notNan),'weights',wbv(notNan).norm,'contourf','antipodal')
% next we plot the WBV form the noisy dataset
for ws = [1 2 3]
wbv = weightedBurgersVec(ebsdN,'windowSize',ws);
nextAxis(1,ws+1)
plot(ebsdN,wbv.norm); hold on
mtexTitle(['WBV norm / box =' num2str(2*ws+1)])
nextAxis(2,ws+1)
notNan = ~isnan(wbv);
plot(wbv(notNan),'weights',wbv(notNan).norm,'contourf','antipodal')
end
mtexColorbar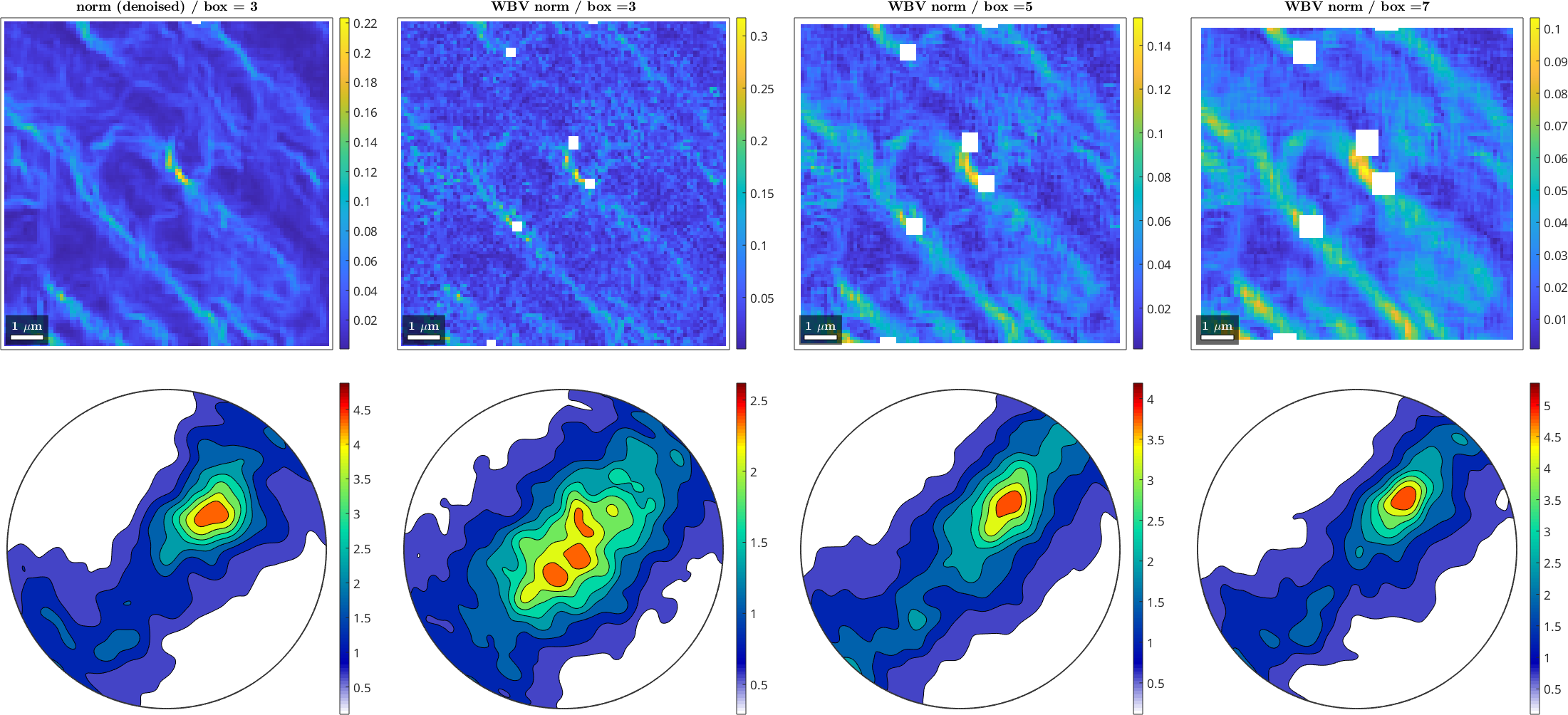
Here we see that there is some difference between the noisy and the denoised data in a 3-by-3 and larger neighborhoods. For larger window sizes, we see that there is of course a loss of detail (and empty spaces appear around unfilled points) and a decrease in the norm of WBV, since high orientation gradients are spread by the larger window size. At the other hand, the distribution of the WBV in the pole figures becomes sharper, because there is less scatter in the WBV over larger areas.
Finally, we look at the weighted Burgers vector computed using the gradient method and the denoised data.
wbv = weightedBurgersVec(ebsd,'gradient');
cK = HSVDirectionKey(specimenSymmetry('1'));
plot(ebsd,cK.direction2color(wbv),'FaceAlpha',wbv.norm/0.2)
mtexTitle('WBV direction / gradient' )
nextAxis(1,2)
notNan = ~isnan(wbv);
plot(wbv(notNan),'weights',wbv(notNan).norm,'antipodal','contourf')
colorbar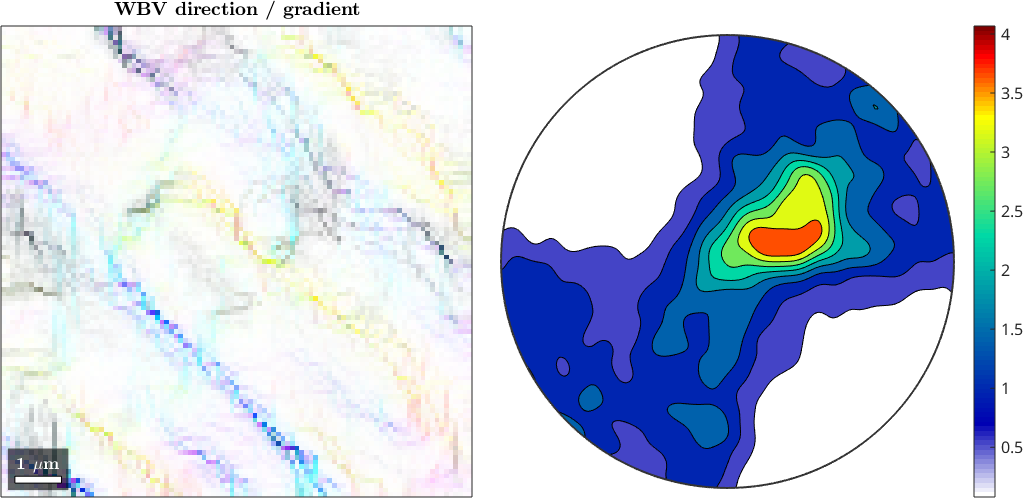
We observe that for denoised data the gradient based method results in a more detailed map.