In this section we explain how to extract specific grain boundaries. Therefore we start by importing some EBSD data and reconstructing the grain structure.
close all;
% import the data
mtexdata forsterite silent
% restrict it to a sub-region of interest.
ebsd = ebsd(inpolygon(ebsd,[5 2 10 5]*10^3));
% and recompute grains
[grains,ebsd.grainId] = calcGrains(ebsd('indexed'),'minPixel',5);
% smooth the grains a bit
grains = smooth(grains,4);
% visualize as a phase map
plot(ebsd)
hold on
plot(grains.boundary,'linewidth',2)
hold off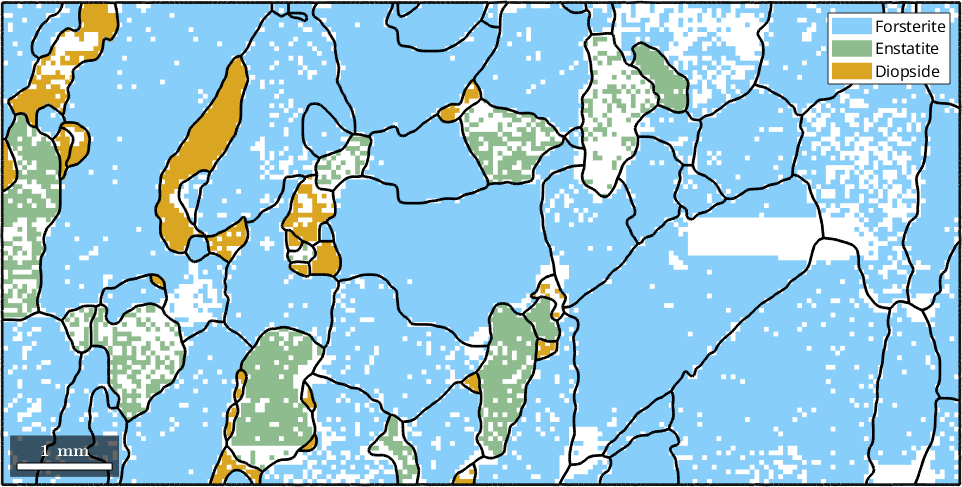
The output of
grains.boundaryans = grainBoundary
Segments length mineral 1 mineral 2
869 27592 µm notIndexed Forsterite
36 1562 µm notIndexed Enstatite
42 1361 µm notIndexed Diopside
1398 56197 µm Forsterite Forsterite
654 26372 µm Forsterite Enstatite
522 20750 µm Forsterite Diopside
35 1296 µm Enstatite Enstatite
134 5802 µm Enstatite Diopside
23 951 µm Diopside Diopsidetells us the number of boundary segments between the different phases. Those segments with phase 'notIndexed' include also those boundary segments where the grains are cut by the scanning boundary. To restrict the grain boundaries to a specific phase transition you shall do
hold on
plot(grains.boundary('Fo','Fo'),'lineColor','blue','micronbar','off','lineWidth',4)
hold off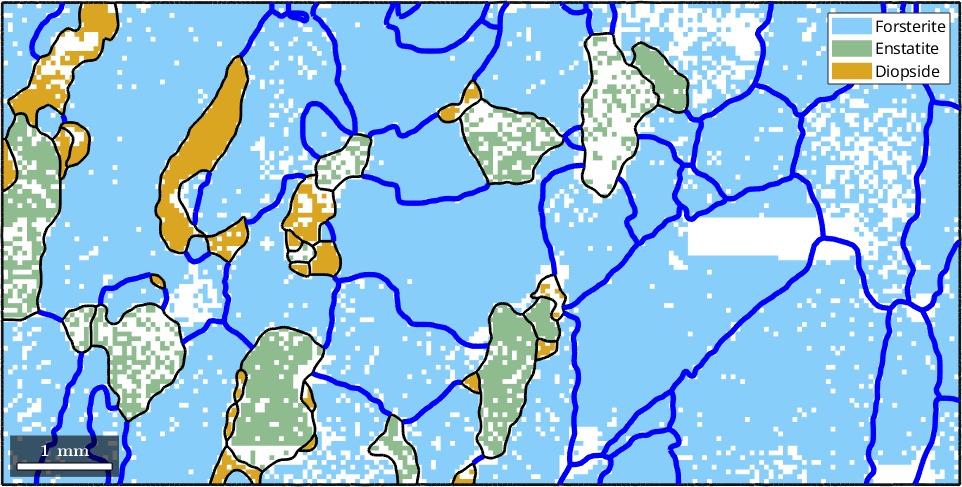
Similarly, we may select all Forsterite to Enstatite boundary segments.
hold on
plot(grains.boundary('Fo','En'),'lineColor','darkgreen','micronbar','off','lineWidth',4)
hold off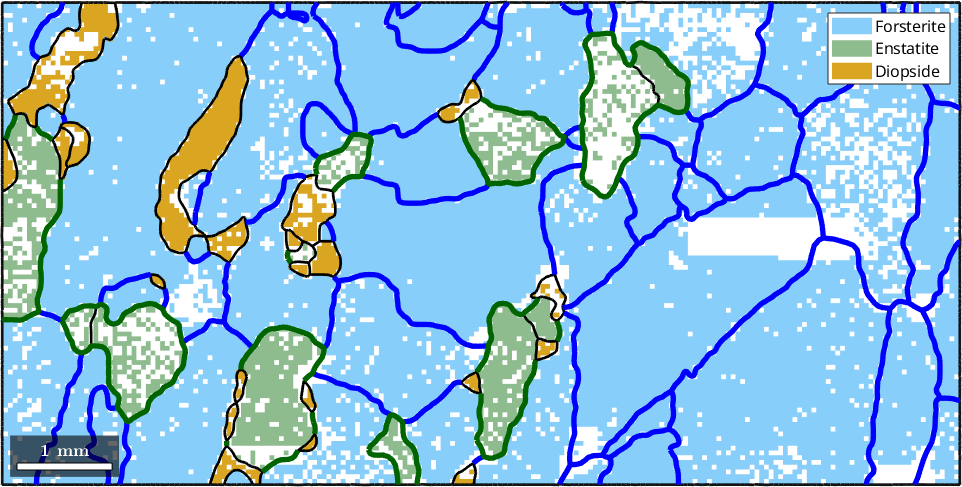
Note, that the order of the phase names matter when considering the corresponding misorientations
grains.boundary('Fo','En').misorientation(1)
grains.boundary('En','Fo').misorientation(1)ans = misorientation (Forsterite → Enstatite)
Bunge Euler angles in degree
phi1 Phi phi2
239.986 50.6875 124.673
ans = misorientation (Enstatite → Forsterite)
Bunge Euler angles in degree
phi1 Phi phi2
55.3267 50.6875 300.014In the fist case the misorientation returned is from Forsterite to Enstatite and in the second case its exactly the inverse
The selection of grain boundaries according to specific misorientations according to twist / tilt character or twinning is explained in linked sections.